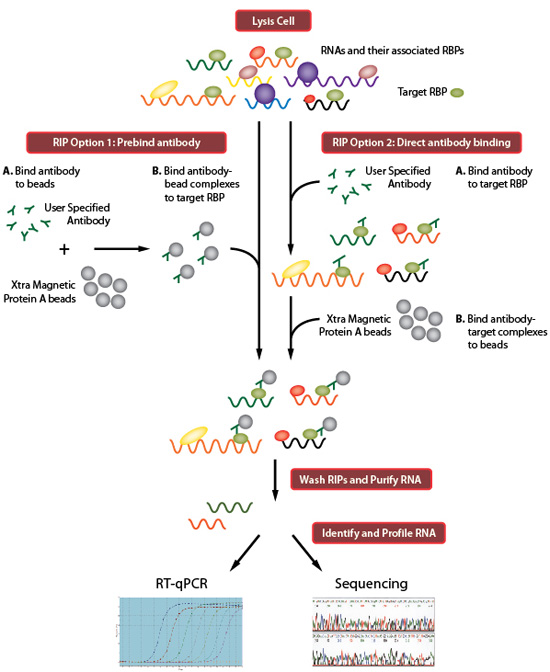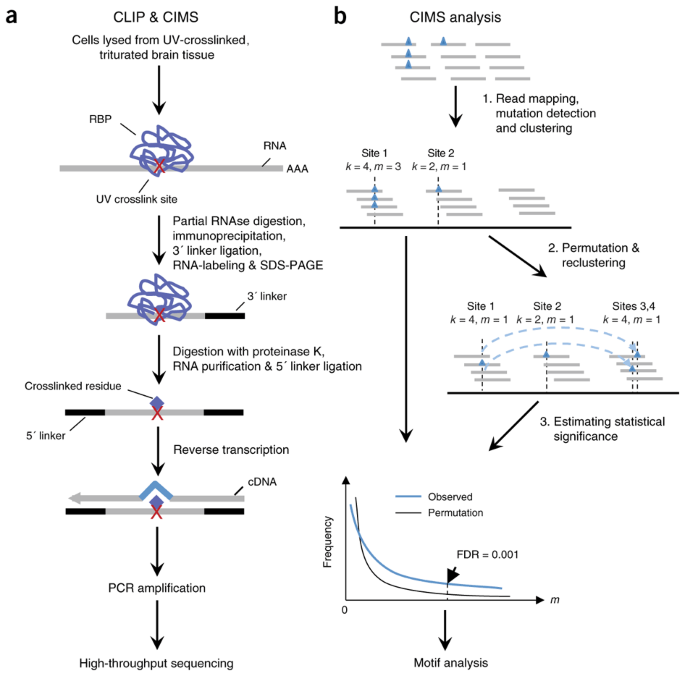
Mapping in vivo protein-RNA interactions at single-nucleotide resolution from HITS-CLIP data | Nature Biotechnology

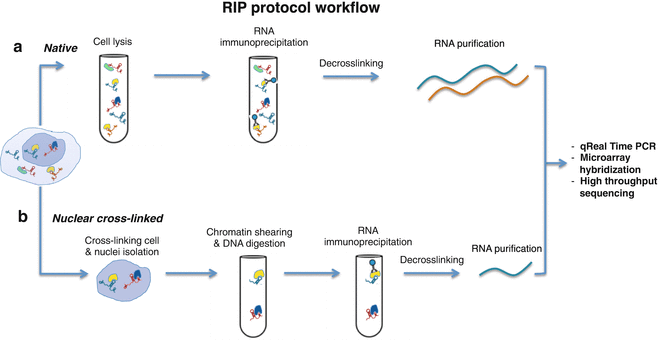
iCLIP - Transcriptome-wide Mapping of Protein-RNA Interactions with Individual Nucleotide Resolution | Protocol
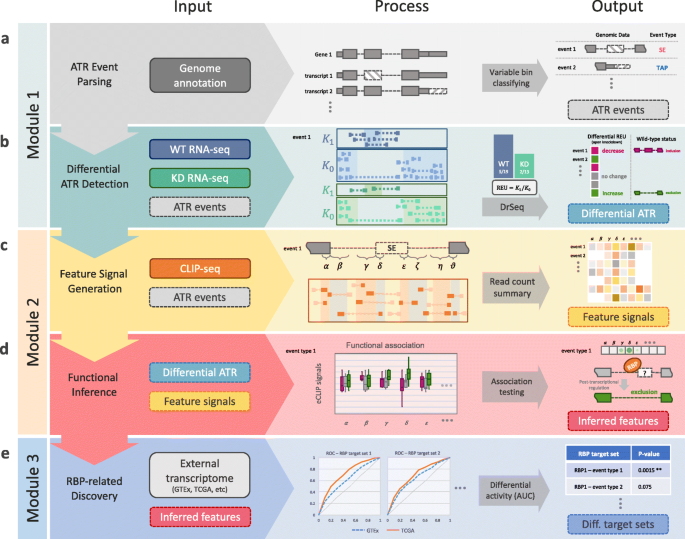
SURF: integrative analysis of a compendium of RNA-seq and CLIP-seq datasets highlights complex governing of alternative transcriptional regulation by RNA-binding proteins | Genome Biology | Full Text
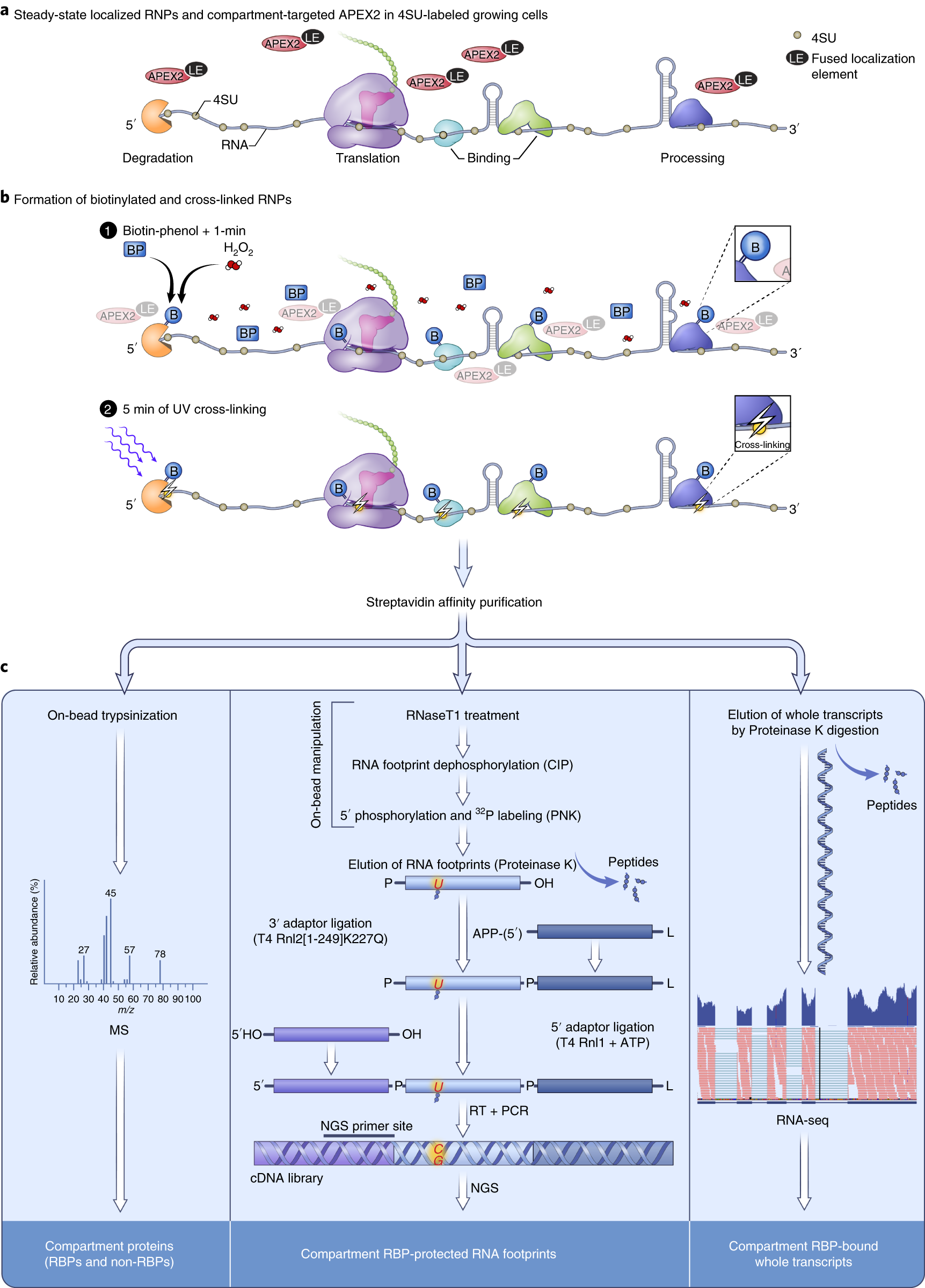
Proximity-CLIP provides a snapshot of protein-occupied RNA elements in subcellular compartments | Nature Methods

Computational approaches for the analysis of RNA–protein interactions: A primer for biologists - Journal of Biological Chemistry

Global Approaches in Studying RNA-Binding Protein Interaction Networks: Trends in Biochemical Sciences

omniCLIP: probabilistic identification of protein-RNA interactions from CLIP-seq data | Genome Biology | Full Text

Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram

Figure 1 | New sequencing methodologies reveal interplay between multiple RNA-binding proteins and their RNAs | SpringerLink

Enhanced CLIP Uncovers IMP Protein-RNA Targets in Human Pluripotent Stem Cells Important for Cell Adhesion and Survival - ScienceDirect

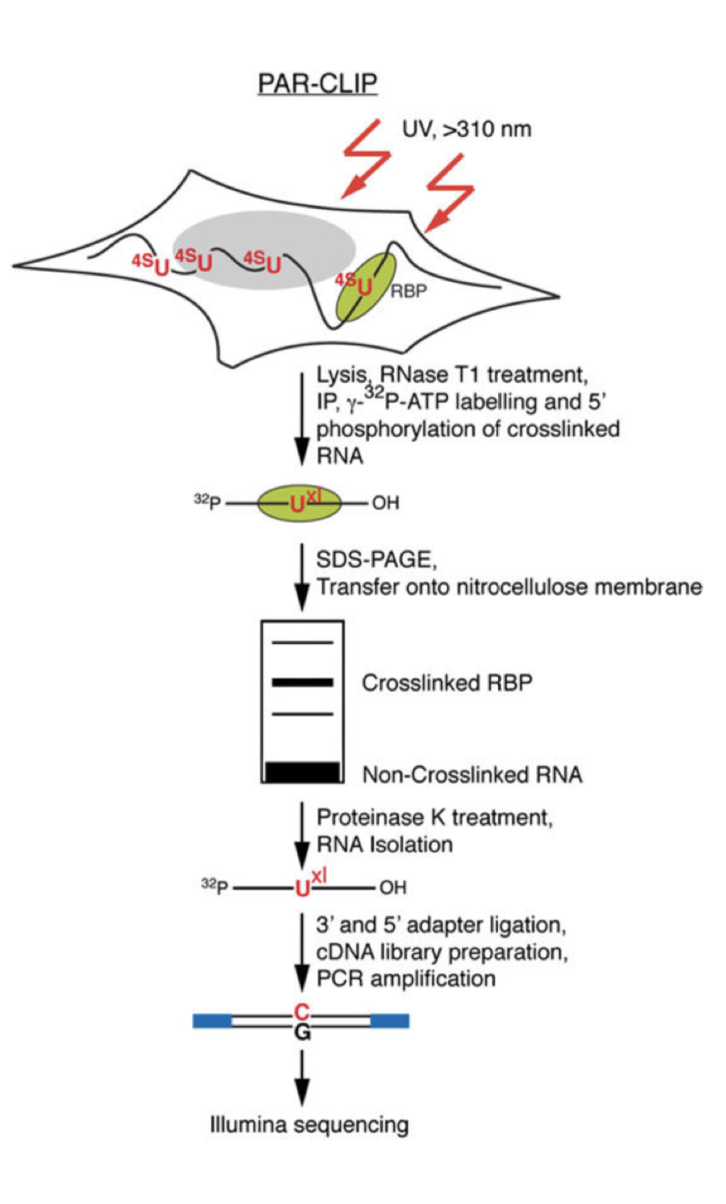
Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

-2.jpg)